Difference between revisions of "Electronic properties of isolated molecules"
Jump to navigation
Jump to search
| Line 42: | Line 42: | ||
*Use the xcrysden software to visualize the relaxation path. | *Use the xcrysden software to visualize the relaxation path. | ||
| − | [[Solution_LAB2_guanine | | + | [[Solution_LAB2_guanine | Solution]] |
===Exercise2=== | ===Exercise2=== | ||
Revision as of 20:41, 18 March 2021
Prev:LabQSM#Lecture 2: DFT simulations of Molecules
Calculation of molecules: Intro
At variance with bulk crystalline systems (fully periodic in 3D), molecules are finite isolated systems, and special care needs to be put in setting up calculations, accordingly.
- scf + nscf: DFT calculations for molecules (or systems with reduced dimensionality in general) follow the same strategies discussed in the previous lectures, where scf calculations are followed by nscf runs (and perhaps by post-processing steps, as discussed below in Electronic properties of isolated molecules#Post processing tools).
- isolation: when localised basis sets are used, systems are usually treated as isolated by default (and care must be used to impose periodicity). Instead, when using plane-waves, periodic boundary conditions (PBC) are automatically enforced and molecule isolation has to be imposed.
- vacuum: As a preliminary step, the unit cell describing an isolated molecule has to be large enough to avoid direct chemicalinteraction (ie chemical bonds) of atoms belonging to different replicas (given PBC, the atoms in one cell are infinitely repeated, giving rise to a lattice of replicas).
- electrostatics: similarly, one also wants to avoid electrostatic interactions (driven in Kohn-Sham DFT by the Hartree potential) among replicas. The presence of long range interactions (dipoles, quadrupoles...) makes convergence wrt vacuum (ie cell size) slow and dedicated techniques should be used. When using Quantum ESPRESSO, consider setting
assume_isolated="mt"(standing for Martyna Tuckerman approach, as from J. Chem. Phys. 110, 2810 (1999)) in theSYSTEMnamelist. - When charged systems are considered the situation is even more critical: in principle the total energy would diverge for a system in PBC, and by default a compensating uniform background is added to the system (though resulting in spurious effects in the properties of the system). In this cases the use of the Martyna-Tuckerman approach is strongly recommended.
- In principle, the exchange-correlation energy and potential would also have a contribution related to the presence of replicas (long range physical contributions are also there, just consider van der Waals interactions as an example). Nevertheless, when using simple approximations to the xc functional (eg LDA, GGA, meta-GGA) long range contributions are not present and so the xc term does not pose any problems.
- Overall, a 10-15 Angstrom of vacuum in between replicas is typically enough to describe neutral molecules with mild dipoles to a decent accuracy. If the Martyna-Tuckerman approach is used, one needs to be very careful with the molecule separation since it needs to be at least as large as the physical dimension of the system in that direction (so for elongated molecules a large region of vacuum is actually needed).
- Importantly: though rules of thumb exist (see above), the amount of vacuum to be included in the system description is a convergence parameter that needs to be tested against the quantities of interested to be calculated.
K-points and Gamma sampling
- The Brillouin zone of a infinitely large cell subject to PBC becomes a single point, Gamma. Because of this, even when using a large but finite cell, K-point sampling should not be used. This should not be relevant in terms of results (if it is, spurious interactions are present), but would result in a waste of computational resources. The so-called Gamma sampling should then be sued.
- Moreover, in view of the Gamma sampling, wave functions can be taken to be real, leading to more memory and computational time savings.
- In Quantum ESPRESSO one needs to set the following:
K_POINTS automatic
1 1 1 0 0 0 # for Gamma sampling using complex wavefunctions
# deprecated, unless for testing or debugging
K_POINTS (gamma) # for proper gamma sampling using real wavefunctions
Post processing tools
Exercises
Exercise1
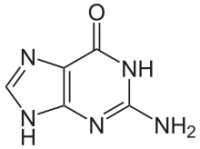
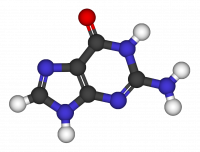
Structural relaxation of the Guanine molecule
- Use a molecular builder to find the initial molecular structure. (Avogadro)
- Classical Force Field Relaxation (using a built-in engine)
- Quantum Mechanics Relaxation refinement (Quantum ESPRESSO) (thr=10^-3)
- Use the xcrysden software to visualize the relaxation path.
Exercise2
Electronic properties of the Guanine molecule
Using the guanine atomic coordinates determined at the previous step, compute:
- HOMO and LUMO frontier orbital energies
- How do they depend on the supercell volume and why?
- Estimate the vacuum level wrt supercell size (pp.x, average.x)
- Repeat the calculation using the “assume_isolated” variable
- Is -HOMO a good estimate of IP ? IP_exp=8.0-8.3 eV